Trap-and-release accelerates study of swimming ciliated cells
J. Mark Meacham and Minji Kim in his lab studied cilia in an acoustic trap that allows them to analyze hundreds of cells in minutes
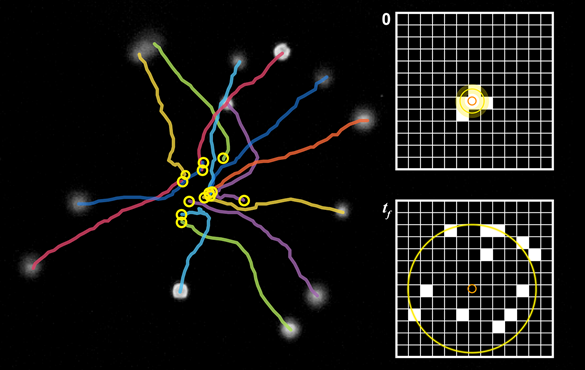
Researchers use ultrasound to streamline measurement of movement
Researchers at Washington University in St. Louis have been studying cilia for years to determine how their dysfunction leads to infertility and other conditions associated with cilia-related diseases. Now, they will be able to perform these studies more rapidly through a new method that uses sound waves to momentarily trap cells propelled by cilia, then releases them to measure their movement as they swim away.
An interdisciplinary team led by J. Mark Meacham, assistant professor of mechanical engineering & materials science in the McKelvey School of Engineering, and students in his lab used an acoustic microfluidic approach that uses ultrasonic standing waves within a small fluid-filled chamber to collect groups of the single-cell green algae cells Chlamydomonas reinhardtii, a model organism for studying human cilia. The so-called acoustic trap takes advantage of material properties of the cell bodies to hold them in place without damaging them. By first collecting the cells, the team can efficiently analyze hundreds of cells in minutes. Results were published in and featured on the inside back cover of the journal Soft Matter in the June 12, 2019, print edition.
"Think of it as a tiny cage made by the ultrasound field," Meacham said. "The cells are trying to find a way to escape but are pushed back by the waves that make up the cage walls. When the walls are removed, they are free to run."
Cilia are tiny hair-like structures in cells that line our lungs, nose, brain and reproductive systems. They are designed to sweep out fluids and microbes to keep people healthy. When they malfunction, infertility, chronic middle-ear infections, water on the brain and other conditions can develop.
Susan Dutcher, professor of genetics and of cell biology and physiology at the School of Medicine and a co-author on the paper, works with C. reinhardtii and hundreds of its genetic variants, or mutants, to study ciliary behavior and dysfunction. Analyzing so many variants using current methods, which manually trace individual cells, would take a very long time, Meacham said.
"It is useful for Dr. Dutcher to rapidly classify her cells based on swimming effectiveness and to choose those that are of most interest for the more laborious and tedious, detailed analysis," Meacham said. "That's what this type of population-based method really helps with, allowing us to analyze a large number of given mutants in a short time."
For this work, the team used three genetic variants of C. reinhardtii cells from Dutcher's lab as models.
Meacham and a doctoral student, Minji Kim, first author on the paper, developed the microfluidic chip, which is small enough that two of them fit on a 1- by 3-inch glass slide. Cells entered and exited through inlet and outlet channels connected to a circular chamber at the center of the device — which is like a large, open holding pen for the cells — before the ultrasound is turned on. Kim and Meacham inserted fluid containing the cells into the device, then activated the ultrasound via a piezoelectric transducer. The ultrasonic waves reflect off of the chamber walls to create pressure wells within the circular chamber, which trap the cells into a group at the chamber's center.
After imaging the cells, the researchers turn off the ultrasound, effectively opening the cage door and allowing the cells to swim off.
"This acoustic trap allows us to do this interesting type of analysis that we couldn't do any other way," Meacham said. "We can trap and release a cell population, analyze it, load up the next population, trap, release, analyze, and load up the next one in a matter of tens of seconds to a minute per sample to get a graded measure of swimming capability for the different cell types."
Analysis of spreading cells is easily automated because swimming starts from a single location, Meacham said. Cells appear as black pixels in successive images of the released cells. The change in shape of the cells is then related to swimming speed.
"We observe them swimming for one to three seconds, then once we have those images, the process of analyzing them is automated," Kim said. "We can get the motility measurement from about 50 cells in an automated way considerably faster than by having to track individual cells."
Ultimately, the team seeks to provide researchers with a tool that categorizes cells based on their movement capability, whether for cataloging C. reinhardtii mutants or for assessing sperm cell motility, Meacham said.
Kim M, Huff E, Bottier M, Dutcher S, Bayly P, Meacham, JM. Acoustic trap-and-release for rapid assessment of cell motility. Soft Matter, 2019, 15, 4266 – 4275, June 12, 2019. DOI: 10.1039/c9sm00184k.
Funding for this research was provided by the National Science Foundation (CMMI-1633971).